AlphaFold
AlphaFold(直譯:阿爾法折疊)是Alphabet旗下Google旗下DeepMind开发的一款蛋白質結構預測程式[1]。該程序被設計為一個深度學習系統[2]。
AlphaFold人工智能有2個主要版本:AlphaFold 1(2018)和AlphaFold 2(2020)。前者使用AlphaFold 1在2018年12月的第13屆CASP(英語:,直譯:蛋白質結構預測的關鍵評估)的排名中第一。該程序特別成功地預測了被競賽組織者評為最困難的目標的最準確結構,其中沒有來自具有部分相似序列的蛋白質的現有模板結構。
蛋白质卷曲折叠會构成三维结构,蛋白质的功能正由其結構決定。了解蛋白質結構有助於開發治療疾病的藥物[3]。DeepMind稱,AlphaFold能在数天内识别蛋白质的形状,而此前學界識別蛋白质形状經常需花費數年時間[4]。2020年11月,在第14届CASP(英語:,直譯:蛋白質結構預測的關鍵評估)競賽中[5],AlphaFold 2(2020)表現良好,中位分数为92.4(满分100分)[6]。它的准确度远远高于其他任何[7] 。
2021年7月15日,AlphaFold 2論文在《自然》雜誌上作為高級訪問出版物與開源軟件和可搜索的物種蛋白質組數據庫一起發表[8][9][10]。
蛋白質折疊問題
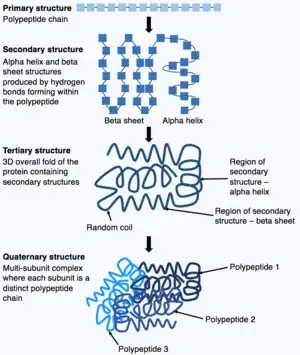
蛋白質由蛋白質一級結構組成,蛋白質折疊的過程中蛋白質會自發折疊形成蛋白質三級結構。蛋白質結構對蛋白質生物學功能至關重要。然而,了解氨基酸序列如何確定蛋白質三級結構極具挑戰性,這被稱為「蛋白質折疊問題」。[11]「蛋白質折疊問題」涉及折疊穩定結構的原子間力熱力學、蛋白質以極快速達到其最終折疊狀態的機制和途徑,以及如何從氨基酸序列預測蛋白質天然結構。[12]
蛋白質結構過去通過諸如X射線晶體學、低溫電子顯微鏡和核磁共振等技術進行實驗確定,這些技術既昂貴又耗時。[11]
過去60年努力只確定了約170,000種蛋白質結構,而所有生命形式中已知蛋白質超過2億種。[13]
如果可以僅從氨基酸序列預測蛋白質結構,將極大地促進科學研究。然而利文索爾佯謬表明,雖蛋白質可在幾毫秒內折疊,但隨機計算所有可能的結構以確定真正的天然結構所需的時間比已知宇宙的年齡要長,這使得預測蛋白質為科學家們構建了生物學中的一項重大挑戰。[11]
多年來,研究人員應用了許多計算方法來解決蛋白質結構預測問題,但除了小而簡單的蛋白質外,它們準確性還遠遠遠沒有接近實驗技術,從而限制了科學研究。
CASP於1994年發起,旨在挑戰科學界做出最好的蛋白質結構預測,結果對於最困難的到2016年的蛋白質發現GDT分數也只能達到100滿分的40分。[13]
算法
DeepMind is known to have trained the program on over 170,000 proteins from a public repository of protein sequences and structures. The program uses a form of attention network, a deep learning technique that focuses on having the AI algorithm identify parts of a larger problem, then piece it together to obtain the overall solution.[2] The overall training was conducted on processing power between 100 and 200 GPUs.[2] Training the system on this hardware took "a few weeks", after which the program would take "a matter of days" to converge for each structure.[14]
AlphaFold 1(2018)
AlphaFold 1 (2018) was built on work developed by various teams in the 2010s, work that looked at the large databanks of related DNA sequences now available from many different organisms (most without known 3D structures), to try to find changes at different residues that appeared to be correlated, even though the residues were not consecutive in the main chain. Such correlations suggest that the residues may be close to each other physically, even though not close in the sequence, allowing a contact map to be estimated. Building on recent work prior to 2018, AlphaFold 1 extended this to estimate a probability distribution for just how close the residues might be likely to be—turning the contact map into a likely distance map. It also used more advanced learning methods than previously to develop the inference. Combining a statistical potential based on this probability distribution with the calculated local free-energy of the configuration, the team was then able to use gradient descent to a solution that best fitted both.[15][16]
More technically, Torrisi et al summarised in 2019 the approach of AlphaFold version 1 as follows:[17]
Central to AlphaFold is a distance map predictor implemented as a very deep residual neural networks with 220 residual blocks processing a representation of dimensionality 64×64×128 – corresponding to input features calculated from two 64 amino acid fragments. Each residual block has three layers including a 3×3 dilated convolutional layer – the blocks cycle through dilation of values 1, 2, 4, and 8. In total the model has 21 million parameters. The network uses a combination of 1D and 2D inputs, including evolutionary profiles from different sources and co-evolution features. Alongside a distance map in the form of a very finely-grained histogram of distances, AlphaFold predicts Φ and Ψ angles for each residue which are used to create the initial predicted 3D structure. The AlphaFold authors concluded that the depth of the model, its large crop size, the large training set of roughly 29,000 proteins, modern Deep Learning techniques, and the richness of information from the predicted histogram of distances helped AlphaFold achieve a high contact map prediction precision.
AlphaFold 2(2020)
The 2020 version of the program (AlphaFold 2, 2020) is significantly different from the original version that won CASP 13 in 2018, according to the team at DeepMind.[18][19]
The DeepMind team had identified that its previous approach, combining local physics with a guide potential derived from pattern recognition, had a tendency to over-account for interactions between residues that were nearby in the sequence compared to interactions between residues further apart along the chain. As a result, AlphaFold 1 had a tendency to prefer models with slightly more secondary structure (alpha helices and beta sheets) than was the case in reality (a form of overfitting).[20]
The software design used in AlphaFold 1 contained a number of modules, each trained separately, that were used to produce the guide potential that was then combined with the physics-based energy potential. AlphaFold 2 replaced this with a system of sub-networks coupled together into a single differentiable end-to-end model, based entirely on pattern recognition, which was trained in an integrated way as a single integrated structure.[19][21] Local physics, in the form of energy refinement based on the AMBER model, is applied only as a final refinement step once the neural network prediction has converged, and only slightly adjusts the predicted structure.[20]
A key part of the 2020 system are two modules, believed to be based on a transformer design, which are used to progressively refine a vector of information for each relationship (or "edge" in graph-theory terminology) between an amino acid residue of the protein and another amino acid residue (these relationships are represented by the array shown in green); and between each amino acid position and each different sequences in the input sequence alignment (these relationships are represented by the array shown in red).[21] Internally these refinement transformations contain layers that have the effect of bringing relevant data together and filtering out irrelevant data (the "attention mechanism") for these relationships, in a context-dependent way, learnt from training data. These transformations are iterated, the updated information output by one step becoming the input of the next, with the sharpened residue/residue information feeding into the update of the residue/sequence information, and then the improved residue/sequence information feeding into the update of the residue/residue information.[21] As the iteration progresses, according to one report, the "attention algorithm ... mimics the way a person might assemble a jigsaw puzzle: first connecting pieces in small clumps—in this case clusters of amino acids—and then searching for ways to join the clumps in a larger whole."[13]
The output of these iterations then informs the final structure prediction module,[21] which also uses transformers,[22] and is itself then iterated. In an example presented by DeepMind, the structure prediction module achieved a correct topology for the target protein on its first iteration, scored as having a GDT_TS of 78, but with a large number (90%) of stereochemical violations – i.e. unphysical bond angles or lengths. With subsequent iterations the number of stereochemical violations fell. By the third iteration the GDT_TS of the prediction was approaching 90, and by the eighth iteration the number of stereochemical violations was approaching zero.[23]
The AlphaFold team stated in November 2020 that they believe AlphaFold can be further developed, with room for further improvements in accuracy.[18]
The training data was originally restricted to single peptide trains. However, the October 2021 update, named AlphaFold-Multimer, included protein complexes in its training data. DeepMind stated this update succeeded about 70% of the time at accurately predicting protein-protein interactions.[24]
競賽
CASP13
In December 2018, DeepMind's AlphaFold placed first in the overall rankings of the 13th Critical Assessment of Techniques for Protein Structure Prediction (CASP).[25][26]
The program was particularly successfully predicting the most accurate structure for targets rated as the most difficult by the competition organisers, where no existing template structures were available from proteins with a partially similar sequence. AlphaFold gave the best prediction for 25 out of 43 protein targets in this class,[26][27][28] achieving a median score of 58.9 on the CASP's global distance test (GDT) score, ahead of 52.5 and 52.4 by the two next best-placed teams,[29] who were also using deep learning to estimate contact distances.[30][31] Overall, across all targets, the program achieved a GDT score of 68.5.[32]
In January 2020, implementations and illustrative code of AlphaFold 1 was released open-source on GitHub.[33][11] but, as stated in the "Read Me" file on that website: "This code can't be used to predict structure of an arbitrary protein sequence. It can be used to predict structure only on the CASP13 dataset (links below). The feature generation code is tightly coupled to our internal infrastructure as well as external tools, hence we are unable to open-source it." Therefore, in essence, the code deposited is not suitable for general use but only for the CASP13 proteins. The company has not announced plans to make their code publicly available as of 5 March 2021.
CASP14
In November 2020, DeepMind's new version, AlphaFold 2, won CASP14.[14][34] Overall, AlphaFold 2 made the best prediction for 88 out of the 97 targets.[35]
On the competition's preferred global distance test (GDT) measure of accuracy, the program achieved a median score of 92.4 (out of 100), meaning that more than half of its predictions were scored at better than 92.4% for having their atoms in more-or-less the right place,[36][37] a level of accuracy reported to be comparable to experimental techniques like X-ray crystallography.[18][38][32] In 2018 AlphaFold 1 had only reached this level of accuracy in two of all of its predictions.[35] 88% of predictions in the 2020 competition had a GDT_TS score of more than 80. On the group of targets classed as the most difficult, AlphaFold 2 achieved a median score of 87.
Measured by the root-mean-square deviation (RMS-D) of the placement of the alpha-carbon atoms of the protein backbone chain, which tends to be dominated by the performance of the worst-fitted outliers, 88% of AlphaFold 2's predictions had an RMS deviation of less than 4 Å for the set of overlapped C-alpha atoms.[35] 76% of predictions achieved better than 3 Å, and 46% had a C-alpha atom RMS accuracy better than 2 Å.,[35] with a median RMS deviation in its predictions of 2.1 Å for a set of overlapped CA atoms.[35] AlphaFold 2 also achieved an accuracy in modelling surface side chains described as "really really extraordinary".
To additionally verify AlphaFold-2 the conference organisers approached four leading experimental groups for structures they were finding particularly challenging and had been unable to determine. In all four cases the three-dimensional models produced by AlphaFold 2 were sufficiently accurate to determine structures of these proteins by molecular replacement. These included target T1100 (Af1503), a small membrane protein studied by experimentalists for ten years.[13]
Of the three structures that AlphaFold 2 had the least success in predicting, two had been obtained by protein NMR methods, which define protein structure directly in aqueous solution, whereas AlphaFold was mostly trained on protein structures in crystals. The third exists in nature as a multidomain complex consisting of 52 identical copies of the same domain, a situation AlphaFold was not programmed to consider. For all targets with a single domain, excluding only one very large protein and the two structures determined by NMR, AlphaFold 2 achieved a GDT_TS score of over 80.
Responses
AlphaFold 2 scoring more than 90 in CASP's global distance test (GDT) is considered a significant achievement in computational biology[13] and great progress towards a decades-old grand challenge of biology.[38] Nobel Prize winner and structural biologist Venki Ramakrishnan called the result "a stunning advance on the protein folding problem",[13] adding that "It has occurred decades before many people in the field would have predicted. It will be exciting to see the many ways in which it will fundamentally change biological research."[14]
Propelled by press releases from CASP and DeepMind,[39][14] AlphaFold 2's success received wide media attention.[40] As well as news pieces in the specialist science press, such as Nature,[38] Science,[13] MIT Technology Review,[2] and New Scientist,[41][42] the story was widely covered by major national newspapers,[43][44][45][46] as well as general news-services and weekly publications, such as Fortune,[47][19] The Economist,[18] Bloomberg,[32] Der Spiegel,[48] and The Spectator.[49] In London The Times made the story its front-page photo lead, with two further pages of inside coverage and an editorial.[50][51] A frequent theme was that ability to predict protein structures accurately based on the constituent amino acid sequence is expected to have a wide variety of benefits in the life sciences space including accelerating advanced drug discovery and enabling better understanding of diseases.[38][52] Writing about the event, the MIT Technology Review noted that the AI had "solved a fifty-year old grand challenge of biology."[2] The same article went on to note that the AI algorithm could "predict the shape of proteins to within the width of an atom."[2]
As summed up by Der Spiegel reservations about this coverage have focussed in two main areas: "There is still a lot to be done" and: "We don't even know how they do it".[53]
Although a 30-minute presentation about AlphaFold 2 was given on the second day of the CASP conference (December 1) by project leader John Jumper,[54] it has been described as "exceedingly high-level, heavy on ideas and insinuations, but almost entirely devoid of detail".[7]Template:Unreliable source Unlike other research groups presenting at CASP14, DeepMind's presentation was not recorded and is not publicly available. DeepMind is expected to publish a scientific paper giving an account of AlphaFold 2 in the proceedings volume of the CASP conference; but it is not known whether it will go beyond what was said in the presentation.
Speaking to El País, researcher Alfonso Valencia said "The most important thing that this advance leaves us is knowing that this problem has a solution, that it is possible to solve it... We only know the result. Google does not provide the software and this is the frustrating part of the achievement because it will not directly benefit science."[46] Nevertheless, as much as Google and DeepMind do release may help other teams develop similar AI systems, an "indirect" benefit.[46] In late 2019 DeepMind released much of the code of the first version of AlphaFold as open source; but only when work was well underway on the much more radical AlphaFold 2. Another option it could take might be to make AlphaFold 2 structure prediction available as an online black-box subscription service. Convergence for a single sequence has been estimated to require on the order of $10,000 worth of wholesale compute time.[55] But this would deny researchers access to the internal states of the system, the chance to learn more qualitatively what gives rise to AlphaFold 2's success, and the potential for new algorithms that could be lighter and more efficient yet still achieve such results. Fears of potential for a lack of transparency by DeepMind have been contrasted with five decades of heavy public investment into the open Protein Data Bank and then also into open DNA sequence repositories, without which the data to train AlphaFold 2 would not have existed.[56][57][58]
Of note, on June 18th, 2021 Demis Hassabis tweeted: "Brief update on some exciting progress on #AlphaFold! We’ve been heads down working flat out on our full methods paper (currently under review) with accompanying open source code and on providing broad free access to AlphaFold for the scientific community. More very soon!"[59]
However it is not yet clear to what extent structure predictions made by AlphaFold 2 will hold up for proteins bound into complexes with other proteins and other molecules.[60] This was not a part of the CASP competition which AlphaFold entered, and not an eventuality it was internally designed to expect. Where structures that AlphaFold 2 did predict were for proteins that had strong interactions either with other copies of themselves, or with other structures, these were the cases where AlphaFold 2's predictions tended to be least refined and least reliable. As a large fraction of the most important biological machines in a cell comprise such complexes, or relate to how protein structures become modified when in contact with other molecules, this is an area that will continue to be the focus of considerable experimental attention.[60]
With so little yet known about the internal patterns that AlphaFold 2 learns to make its predictions, it is not yet clear to what extent the program may be impaired in its ability to identify novel folds, if such folds are not well represented in the existing protein structures known in structure databases.[61][60] It is also not well known the extent to which protein structures in such databases, overwhelmingly of proteins that it has been possible to crystallise to X-ray, are representative of typical proteins that have not yet been crystallised. And it is also unclear how representative the frozen protein structures in crystals are of the dynamic structures found in the cells in vivo. AlphaFold 2's difficulties with structures obtained by protein NMR methods may not be a good sign.
On its potential as a tool for drug discovery, Stephen Curry notes that while the resolution of AlphaFold 2's structures may be very good, the accuracy with which binding sites are modelled needs to be even higher: typically molecular docking studies require the atomic positions to be accurate within a 0.3 Å margin, but the predicted protein structure only have at best an RMSD of 0.9 Å for all atoms. So AlphaFold 2's structures may only be a limited help in such contexts.[61][60] Moreover, according to Science columnist Derek Lowe, because the prediction of small-molecule binding even then is still not very good, computational prediction of drug targets is simply not in a position to take over as the "backbone" of corporate drug discovery—so "protein structure determination simply isn’t a rate-limiting step in drug discovery in general".[62] It has also been noted that even with a structure for a protein, to then understand how it functions, what it does, and how that fits within wider biological processes can still be very challenging.[63] Nevertheless, if better knowledge of protein structure could lead to better understanding of individual disease mechanisms and ultimately to better drug targets, or better understanding of the differences between human and animal models, ultimately that could lead to improvements.[64]
Also, because AlphaFold processes protein-only sequences by design, other associated biomolecules are not considered. On the impact of absent metals, co-factors and, most visibly, co- and post-translational modifications such as protein glycosylation from AlphaFold models, Elisa Fadda (Maynooth University, Ireland) and Jon Agirre (University of York, UK) highlighted the need for scientists to check databases such as UniProt-KB for likely missing components, as these can play an important role not just in folding but in protein function.[65] However, the authors highlighted that many AlphaFold models were accurate enough to allow for the introduction of post-predictional modifications.[65]
Finally, some have noted that even a perfect answer to the protein prediction problem would still leave questions about the protein folding problem—understanding in detail how the folding process actually occurs in nature (and how sometimes they can also misfold).[66]
But even with such caveats, AlphaFold 2 was described as a huge technical step forward and intellectual achievement.[67][68]
AlphaFold蛋白質結構數據庫
AlphaFold蛋白質結構數據庫於2021年7月22日啟動,這是AlphaFold和歐洲分子生物學實驗室的歐洲生物信息研究所的共同努力。AlphaFold提供對超過2億個蛋白質結構預測的開放訪問,以加速科學研究。在啟動時,該數據庫包含人類和20種模式生物的幾乎完整UniProt蛋白質組的AlphaFold預測蛋白質結構模型,總計超過365,000種蛋白質(該數據庫不包括少於16個或多於2700個氨基酸殘基蛋白質[69],但對人類而言,殘基蛋白質可在文件中獲得。[70])。
AlphaFold目標是覆蓋UniRef90中1億個蛋白質大部分集合。截至2022年5月15日,已有992,316個可用。[71]
應用
AlphaFold已被用於預測SARS-CoV-2(COVID-19的病原體)的蛋白質結構。 這些蛋白質的結構在2020年初有待實驗檢測[72]。在將結果發佈到更大的研究界之前,英國弗朗西斯·克里克研究所(Francis Crick Institute)的科學家們對結果進行了檢查。該團隊還證實了對實驗確定的SARS-CoV-2刺突蛋白的準確預測,該蛋白在國際開放存取數據庫蛋白質資料庫(Protein Data Bank)中共享,然後發布了計算確定的未充分研究的蛋白質分子的結構[73]。
参考文献
- . Deepmind. [2020-11-30]. (原始内容存档于2021-01-19).
- . MIT Technology Review. [2020-11-30]. (原始内容存档于2021-08-28) (英语).
- . 第一財經.
- . 金融時報中文網. [2020-12-03]. (原始内容存档于2020-12-22).
- Shead, Sam. . CNBC. 2020-11-30 [2020-11-30]. (原始内容存档于2021-01-28) (英语).
- . 科技日报. [2020-12-03]. (原始内容存档于2020-12-05).
- . MIT Technology Review. [2020-11-30]. (原始内容存档于2021-08-28) (英语).
- Jumper, John; Evans, Richard; Pritzel, Alexander; Green, Tim; Figurnov, Michael; Ronneberger, Olaf; Tunyasuvunakool, Kathryn; Bates, Russ; Žídek, Augustin; Potapenko, Anna; Bridgland, Alex; Meyer, Clemens; Kohl, Simon A A; Ballard, Andrew J; Cowie, Andrew; Romera-Paredes, Bernardino; Nikolov, Stanislav; Jain, Rishub; Adler, Jonas; Back, Trevor; Petersen, Stig; Reiman, David; Clancy, Ellen; Zielinski, Michal; Steinegger, Martin; Pacholska, Michalina; Berghammer, Tamas; Bodenstein, Sebastian; Silver, David; Vinyals, Oriol; Senior, Andrew W; Kavukcuoglu, Koray; Kohli, Pushmeet; Hassabis, Demis. . Nature. 2021-07-15, 596 (7873): 583–589. PMC 8371605
 . PMID 34265844. doi:10.1038/s41586-021-03819-2
. PMID 34265844. doi:10.1038/s41586-021-03819-2  (英语).
(英语). - . GitHub. [2021-07-24]. (原始内容存档于2021-07-23) (英语).
- . alphafold.ebi.ac.uk. [2021-07-24]. (原始内容存档于2021-07-24).
- . Deepmind. [2020-11-30]. (原始内容存档于2022-03-07).
- Ken A. Dill, S. Banu Ozkan, M. Scott Shell, and Thomas R. Weikl. . Annual Review of Biophysics. 2008, 37: 289–316. PMC 2443096
 . PMID 18573083. doi:10.1146/annurev.biophys.37.092707.153558.
. PMID 18573083. doi:10.1146/annurev.biophys.37.092707.153558. - Robert F. Service, 'The game has changed.' AI triumphs at solving protein structures (页面存档备份,存于), Science, 30 November 2020
- . Deepmind. [2020-11-30]. (原始内容存档于2020-11-30).
- Mohammed AlQuraishi (May 2019), AlphaFold at CASP13 (页面存档备份,存于), Bioinformatics, 35(22), 4862–4865 doi:10.1093/bioinformatics/btz422. See also Mohammed AlQuraishi (December 9, 2018), AlphaFold @ CASP13: "What just happened?" (页面存档备份,存于) (blog post).
Mohammed AlQuraishi (15 January 2020), A watershed moment for protein structure prediction (页面存档备份,存于), Nature 577, 627–628 doi:10.1038/d41586-019-03951-0 - AlphaFold: Machine learning for protein structure prediction (页面存档备份,存于), Foldit, 31 January 2020
- Torrisi, Mirko et al. (22 Jan. 2020), Deep learning methods in protein structure prediction (页面存档备份,存于). Computational and Structural Biotechnology Journal vol. 18 1301–1310. doi:10.1016/j.csbj.2019.12.011 (CC-BY-4.0)
- . The Economist. 2020-11-30 [2020-11-30]. ISSN 0013-0613. (原始内容存档于2020-12-03).
- Jeremy Kahn, Lessons from DeepMind's breakthrough in protein-folding A.I. (页面存档备份,存于), Fortune, 1 December 2020
- John Jumper et al., conference abstract (December 2020)
- See block diagram. Also John Jumper et al. (1 December 2020), AlphaFold 2 presentation (页面存档备份,存于), slide 10
- The structure module is stated to use a "3-d equivariant transformer architecture" (John Jumper et al. (1 December 2020), AlphaFold 2 presentation (页面存档备份,存于), slide 12).
One design for a transformer network with SE(3)-equivariance was proposed in Fabian Fuchs et al SE(3)-Transformers: 3D Roto-Translation Equivariant Attention Networks (页面存档备份,存于), NeurIPS 2020; also website (页面存档备份,存于). It is not known how similar this may or may not be to what was used in AlphaFold.
See also the blog post (页面存档备份,存于) by AlQuaraishi on this, or the more detailed post (页面存档备份,存于) by Fabian Fuchs - John Jumper et al. (1 December 2020), AlphaFold 2 presentation (页面存档备份,存于), slides 12 to 20
- Callaway, Ewen. . Nature. 2022-04-13, 604 (7905): 234–238 [2022-04-15]. doi:10.1038/d41586-022-00997-5. (原始内容存档于2022-07-26) (英语).
- Group performance based on combined z-scores (页面存档备份,存于), CASP 13, December 2018. (AlphaFold = Team 043: A7D)
- Sample, Ian. . The Guardian. 2018-12-02 [2020-11-30]. (原始内容存档于2019-07-18).
- . Deepmind. [2020-11-30].
- Singh, Arunima. . Nature Methods. 2020, 17 (3): 249. ISSN 1548-7105. PMID 32132733. S2CID 212403708. doi:10.1038/s41592-020-0779-y
 (英语).
(英语). - See CASP 13 data tables (页面存档备份,存于) for 043 A7D, 322 Zhang, and 089 MULTICOM
- Wei Zheng et al,Deep-learning contact-map guided protein structure prediction in CASP13 (页面存档备份,存于), Proteins: Structure, Function, and Bioinformatics, 87(12) 1149–1164 doi:10.1002/prot.25792; and slides (页面存档备份,存于)
- Hou, Jie; Wu, Tianqi; Cao, Renzhi; Cheng, Jianlin. . Proteins: Structure, Function, and Bioinformatics (Wiley). 2019-04-25, 87 (12): 1165–1178. ISSN 0887-3585. PMC 6800999
 . PMID 30985027. bioRxiv 10.1101/552422
. PMID 30985027. bioRxiv 10.1101/552422  . doi:10.1002/prot.25697.
. doi:10.1002/prot.25697. - . Bloomberg.com. 2020-11-30 [2020-11-30]. (原始内容存档于2022-04-05) (英语).
- . GitHub. [2020-11-30]. (原始内容存档于2022-02-01) (英语).
- . MIT Technology Review. [2020-11-30]. (原始内容存档于2021-08-28) (英语).
- Mohammed AlQuraishi, CASP14 scores just came out and they’re astounding (页面存档备份,存于), Twitter, 30 November 2020.
- For the GDT_TS measure used, each atom in the prediction scores a quarter of a point if it is within 8 Å(0.80 nm) of the experimental position; half a point if it is within 4 Å, three-quarters of a point if it is within 2 Å, and a whole point if it is within 1 Å.
- To achieve a GDT_TS score of 92.5, mathematically at least 70% of the structure must be accurate to within 1 Å, and at least 85% must be accurate to within 2 Å.
- Callaway, Ewen. . Nature. 2020-11-30, 588 (7837): 203–204. Bibcode:2020Natur.588..203C. PMID 33257889. doi:10.1038/d41586-020-03348-4
 (英语).
(英语). - Artificial intelligence solution to a 50-year-old science challenge could ‘revolutionise’ medical research (页面存档备份,存于) (press release), CASP organising committee, 30 November 2020
- Brigitte Nerlich, Protein folding and science communication: Between hype and humility (页面存档备份,存于), University of Nottingham blog, 4 December 2020
- Michael Le Page, DeepMind's AI biologist can decipher secrets of the machinery of life (页面存档备份,存于), New Scientist, 30 November 2020
- The predictions of DeepMind’s latest AI could revolutionise medicine (页面存档备份,存于), New Scientist, 2 December 2020
- Cade Metz, London A.I. Lab Claims Breakthrough That Could Accelerate Drug Discovery (页面存档备份,存于), New York Times, 30 November 2020
- Ian Sample,DeepMind AI cracks 50-year-old problem of protein folding (页面存档备份,存于), The Guardian, 30 November 2020
- Lizzie Roberts, 'Once in a generation advance' as Google AI researchers crack 50-year-old biological challenge (页面存档备份,存于). Daily Telegraph, 30 November 2020
- Nuño Dominguez, La inteligencia artificial arrasa en uno de los problemas más importantes de la biología (页面存档备份,存于) (Artificial intelligence takes out one of the most important problems in biology), El País, 2 December 2020
- Jeremy Kahn, In a major scientific breakthrough, A.I. predicts the exact shape of proteins (页面存档备份,存于), Fortune, 30 November 2020
- Julia Merlot, Forscher hoffen auf Durchbruch für die Medikamentenforschung (页面存档备份,存于) (Researchers hope for a breakthrough for drug research), Der Spiegel, 2 December 2020
- Bissan Al-Lazikani, The solving of a biological mystery (页面存档备份,存于), The Spectator, 1 December 2020
- Tom Whipple, "Deepmind computer solves new puzzle: life", The Times, 1 December 2020. front page image (页面存档备份,存于), via Twitter.
- Tom Whipple, Deepmind finds biology’s ‘holy grail’ with answer to protein problem (页面存档备份,存于), The Times (online), 30 November 2020.
In all science editor Tom Whipple wrote six articles on the subject for The Times on the day the news broke. (thread (页面存档备份,存于)). - Tim Hubbard, The secret of life, part 2: the solution of the protein folding problem. (页面存档备份,存于), medium.com, 30 November 2020
- Christian Stöcker, Google greift nach dem Leben selbst (页面存档备份,存于) (Google is reaching for life itself), Der Spiegel, 6 December 2020
- John Jumper et al. (1 December 2020), AlphaFold 2 (页面存档备份,存于). Presentation given at CASP 14.
- Carlos Outeiral, CASP14: what Google DeepMind’s AlphaFold 2 really achieved, and what it means for protein folding, biology and bioinformatics (页面存档备份,存于), Oxford Protein Informatics Group. (3 December)
- Aled Edwards, The AlphaFold2 success: It took a village (页面存档备份,存于), via medium.com, 5 December 2020
- David Briggs, If Google’s Alphafold2 really has solved the protein folding problem, they need to show their working (页面存档备份,存于), The Skeptic, 4 December 2020
- The Guardian view on DeepMind’s brain: the shape of things to come (页面存档备份,存于), The Guardian, 6 December 2020
- Demis Hassabis, "Brief update on some exciting progress on #AlphaFold!" (页面存档备份,存于) (tweet), via twitter, 18 June 2021
- Tom Ireland, How will AlphaFold change bioscience research? (页面存档备份,存于), The Biologist, 4 December 2020
- Stephen Curry, No, DeepMind has not solved protein folding (页面存档备份,存于), Reciprocal Space (blog), 2 December 2020
- Derek Lowe, In the Pipeline: What’s Crucial And What Isn’t (页面存档备份,存于), Science Translational Medicine, 25 September 2019
- Philip Ball, Behind the Screens of AlphaFold (页面存档备份,存于), Chemistry World, 9 December 2020. See also tweets (页面存档备份,存于), 1 December
- Derek Lowe, In the Pipeline: The Big Problems (页面存档备份,存于), Science Translational Medicine, 1 December 2020
- Bagdonas, Haroldas; Fogarty, Carl A.; Fadda, Elisa; Agirre, Jon. . Nature Structural & Molecular Biology. 2021-10-29, 28 (11): 869–870 [2022-07-29]. ISSN 1545-9985. PMID 34716446. S2CID 240228913. doi:10.1038/s41594-021-00680-9. (原始内容存档于2022-06-23) (英语).
- e.g. Greg Bowman, Protein folding and related problems remain unsolved despite AlphaFold's advance (页面存档备份,存于), Folding@home blog, 8 December 2020
- Cristina Sáez, El último avance fundamental de la biología se basa en la investigación de un científico español (页面存档备份,存于), La Vanguardia, 2 December 2020. (Alfonso Valencia overall view)
- Zero Gravitas and Jacky Liang, DeepMind’s AlphaFold 2—An Impressive Advance With Hyperbolic Coverage (页面存档备份,存于), Skynet today (blog), Stanford, 9 December 2020
- . alphafold.ebi.ac.uk. [2021-07-29]. (原始内容存档于2022-07-29).
- . alphafold.ebi.ac.uk. [2021-07-27]. (原始内容存档于2022-07-29).
- . www.alphafold.ebi.ac.uk. [2022-07-29]. (原始内容存档于2022-08-02).
- . Wired. [2020-12-01]. ISSN 1059-1028. (原始内容存档于2022-04-23) (美国英语).
- . Deepmind. [2020-12-01]. (原始内容存档于2022-03-25).
外部链接
AlphaFold(2018年)
- AlphaFold: The making of a scientific breakthrough (页面存档备份,存于), DeepMind, via YouTube.
- Senior, Andrew. . Institute for Protein Design. 2019-08-23 [2020-12-03]. (原始内容存档于2020-12-19) –YouTube.
- GitHub上的AlphaFold code used at CASP13
- GitHub上的Open source community implementationTemplate:SpndProSPr
AlphaFold 2(2020年)
- GitHub上的AlphaFold v2.1 code and links to model
- Open access to protein structure predictions for the human proteome and 20 other key organisms (页面存档备份,存于) at 欧洲生物信息研究所(EMBL-EBI)
- CASP 14 (页面存档备份,存于) website
- AlphaFold: The making of a scientific breakthrough (页面存档备份,存于), DeepMind, via YouTube.
- ColabFold (页面存档备份,存于) (Mirdita, Milot; Ovchinnikov, Sergey; Steinegger, Martin. . 2021-08-15. bioRxiv 10.1101/2021.08.15.456425
 (英语). ), version (页面存档备份,存于) for homooligomeric prediction and complexes (页面存档备份,存于)
(英语). ), version (页面存档备份,存于) for homooligomeric prediction and complexes (页面存档备份,存于)